Blog Categories
RNA-Seq Case Study: Genes Regulated by p53-SET Interplay
The oncoprotein SET can bind to p53 and inhibits p53 transcriptional activity in unstressed cells. To identify the genes regulated by SET in a p53-dependent manner, the endogenous SET was depleted by siRNA in control or p53 knockout U2OS cells. We use BxGenomics DEG and Venn Diagram tools to find genes regulated by p53-SET.
First we use the study GEO ID GSE83635 to find the study.
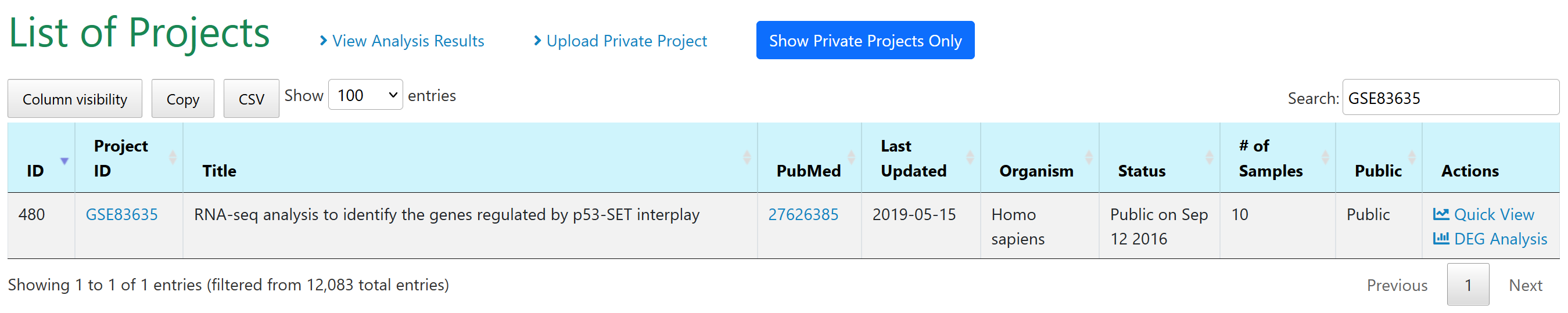
Next we can choose "DEG Analysis" under Actions column to perform three comparisons.
- WT_si-SET.vs.WT_siCtrl. In normal cells, genes that are induced by SET siRNA depletion are upregulated in this comparison.
- WT_si-SET.vs.KO_si-SET. When SET is depleted, genes whose activation are abolished by p53 knockdown are upregulated in this comparison.
- WT_siCtrl.vs.KO_siCtrl. When SET is normal, genes whose expression are abolished by p53 knockdown are upregulated in this comparison.
In the analysis results, when we use Venn Diagram tool to check the up-regulated genes from the three comparisons, we can see there are a lot of overlap between the three comparisons.
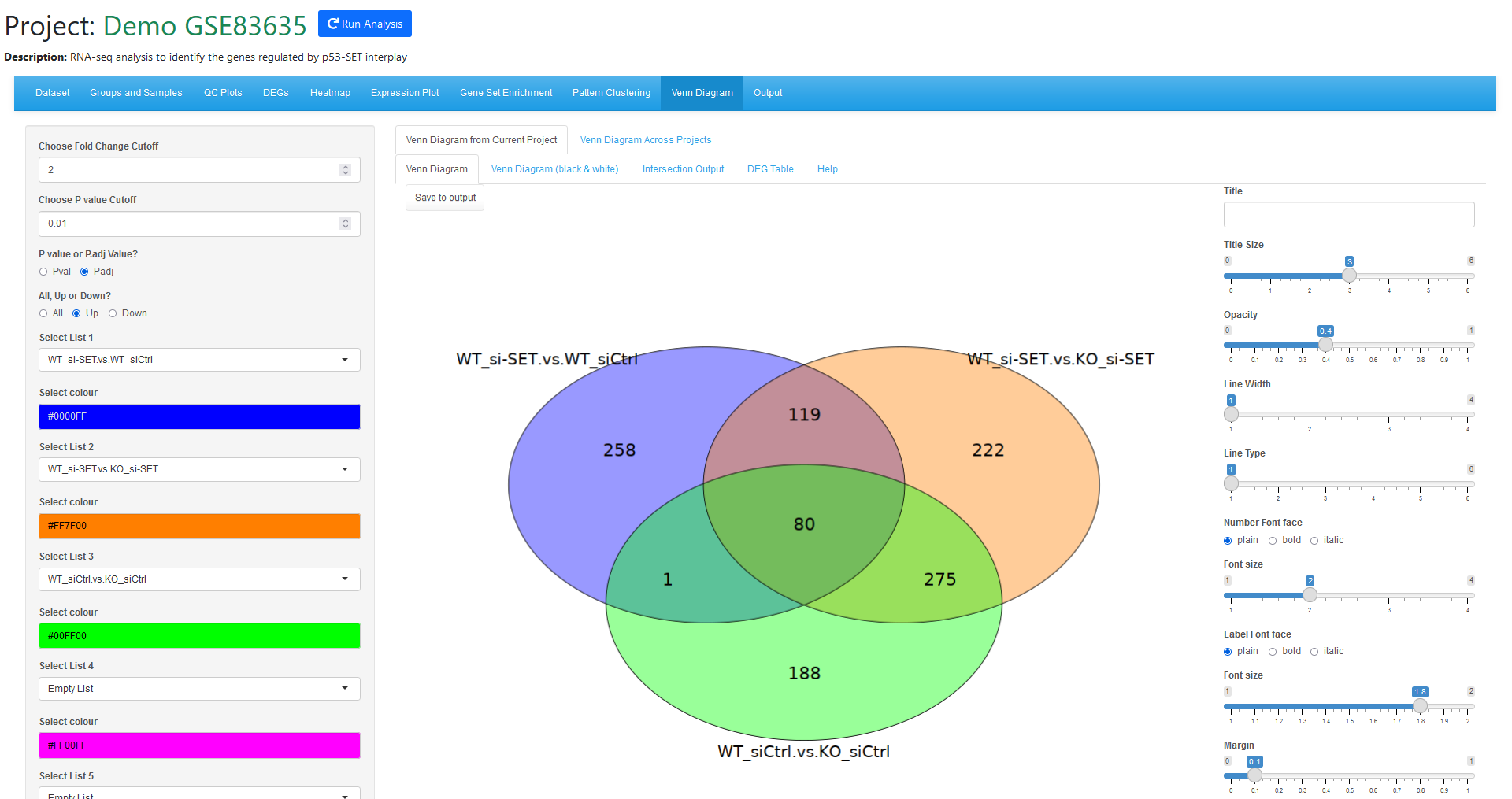
We can get the 80 genes from the intersection output tab from the Venn Diagram tool, the use this gene list in the heatmap tool with "Upload Genes" option. We added SET genes to the list so we can view SET depletion by siRNA.
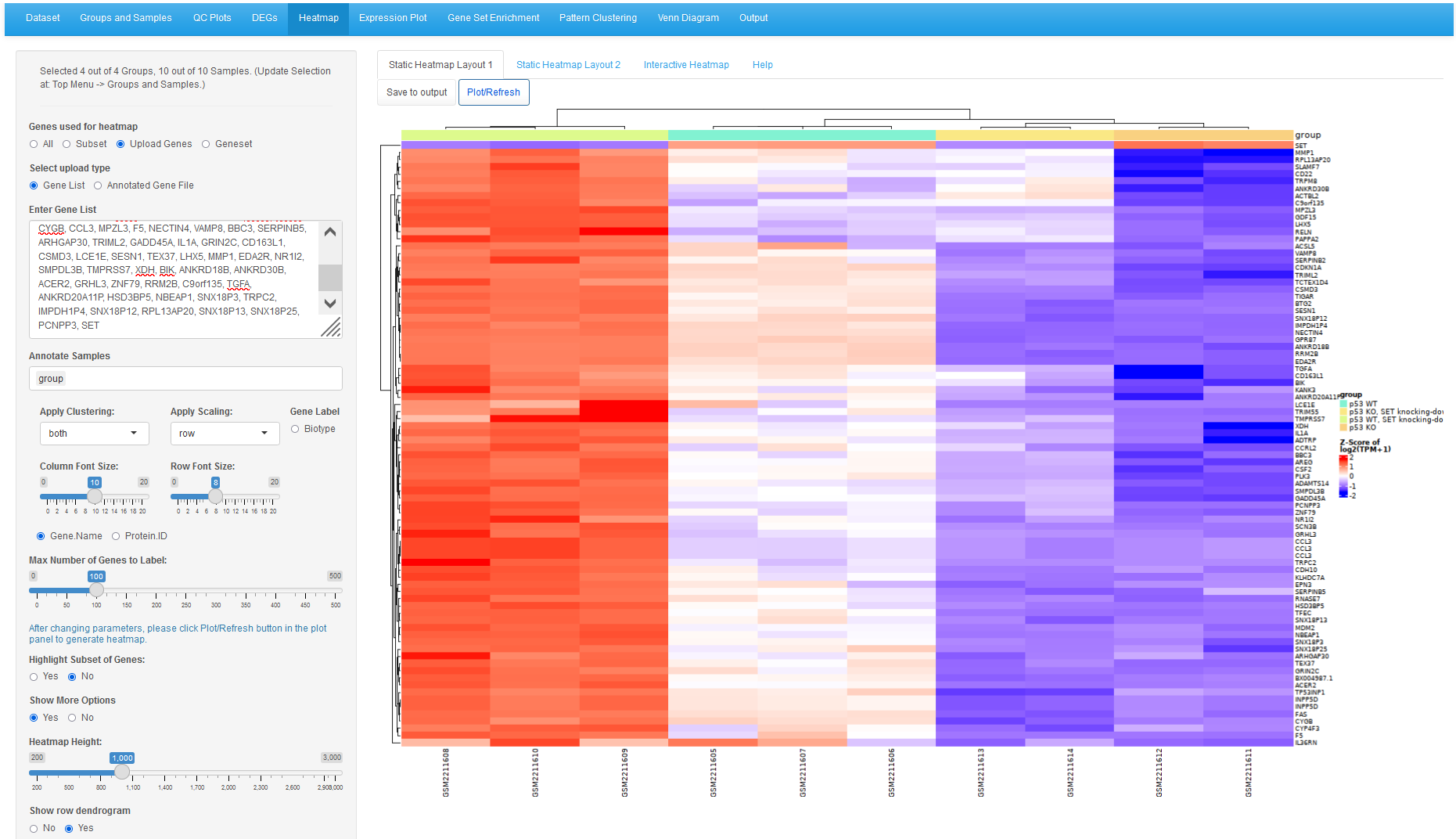
SET gene is the first gene in the heatmap, siRNA depletion is clear. The rest are the 80 genes from the Venn diagram intersection. It can be seen that these 80 genes are most likely regulated by p53-SET interplay. These genes have basal level of expression in p53 WT cells, and are highly expressed when SET is depleted by siRNA. However, in p53 KO cells, these genes are not expressed, and depleting PET doesn't induce these genes when p53 is knocked out.
Many of the genes found by this analysis (e.g. TGFA, FAS, BBC3, BTG2) are known p53 target genes. Indeed, if we check upregualgted genes in WT_si-SET.vs.WT_siCtrl, p53_pathway come up.